Nápady Color Atoms Pymol Výborně
Nápady Color Atoms Pymol Výborně. Pymol can deduce bonds from the pdb structure file, even if the conect records are missing Usually, people have to draw the protein structure according to folding, titration or relaxation experiments; The c alpha and the amide bond's c and n atom. When the script is run in pymol, hydrogen atoms are removed and the exact colors are defined with rgb color values. To retrieve the color for all residues in a selection, you can iterate over it from the pymol …
Nejlepší Tips
To retrieve the color for all residues in a selection, you can iterate over it from the pymol … A hash table is used to map amino acids to a list of ordered pairs of atoms with their color according to their position in the side chain and which atoms they are bound to (figure 1). Color by atom type from a script. Robert campbell has a color_b.py python script on his pymol homepage that you can use.I'm not sure how you imagine it should look if only the n part of this representation should be colored.
See section color for this. When the script is run in pymol, hydrogen atoms are removed and the exact colors are defined with rgb color values. Description spectrum colors atoms with a spectrum of colors based on an atomic property. See section color for this. Pymol api cmd.color(string color, string selection, int quiet) see also color_deep, set_color, recolor example color cyan color yellow, chain a.
I'm not sure how you imagine it should look if only the n part of this representation should be colored. Pymol can deduce bonds from the pdb structure file, even if the conect records are missing A hash table is used to map amino acids to a list of ordered pairs of atoms with their color according to their position in the side chain and which atoms they are bound to (figure 1). Description spectrum colors atoms with a spectrum of colors based on an atomic property. To retrieve the color for all residues in a selection, you can iterate over it from the pymol … Robert campbell has a color_b.py python script on his pymol homepage that you can use.. To retrieve the color for all residues in a selection, you can iterate over it from the pymol …

Pymol api cmd.color(string color, string selection, int quiet) see also color_deep, set_color, recolor example color cyan color yellow, chain a. Robert campbell has a color_b.py python script on his pymol homepage that you can use. A hash table is used to map amino acids to a list of ordered pairs of atoms with their color according to their position in the side chain and which atoms they are bound to (figure 1). Description spectrum colors atoms with a spectrum of colors based on an atomic property. When the script is run in pymol, hydrogen atoms are removed and the exact colors are defined with rgb color values. I'm not sure how you imagine it should look if only the n part of this representation should be colored.

Robert campbell has a color_b.py python script on his pymol homepage that you can use... A hash table is used to map amino acids to a list of ordered pairs of atoms with their color according to their position in the side chain and which atoms they are bound to (figure 1). I'm not sure how you imagine it should look if only the n part of this representation should be colored. Pymol can deduce bonds from the pdb structure file, even if the conect records are missing
Pymol can deduce bonds from the pdb structure file, even if the conect records are missing The c alpha and the amide bond's c and n atom. Pymol can deduce bonds from the pdb structure file, even if the conect records are missing A hash table is used to map amino acids to a list of ordered pairs of atoms with their color according to their position in the side chain and which atoms they are bound to (figure 1). To retrieve the color for all residues in a selection, you can iterate over it from the pymol … Description spectrum colors atoms with a spectrum of colors based on an atomic property. See section color for this. Usually, people have to draw the protein structure according to folding, titration or relaxation experiments;

Pymol api cmd.color(string color, string selection, int quiet) see also color_deep, set_color, recolor example color cyan color yellow, chain a. . Usually, people have to draw the protein structure according to folding, titration or relaxation experiments;

Description spectrum colors atoms with a spectrum of colors based on an atomic property. When the script is run in pymol, hydrogen atoms are removed and the exact colors are defined with rgb color values. I'm not sure how you imagine it should look if only the n part of this representation should be colored. Usually, people have to draw the protein structure according to folding, titration or relaxation experiments; Pymol api cmd.color(string color, string selection, int quiet) see also color_deep, set_color, recolor example color cyan color yellow, chain a. To retrieve the color for all residues in a selection, you can iterate over it from the pymol … Robert campbell has a color_b.py python script on his pymol homepage that you can use. Color by atom type from a script. The c alpha and the amide bond's c and n atom.

Pymol api cmd.color(string color, string selection, int quiet) see also color_deep, set_color, recolor example color cyan color yellow, chain a. I'm not sure how you imagine it should look if only the n part of this representation should be colored.
See section color for this.. Robert campbell has a color_b.py python script on his pymol homepage that you can use. When the script is run in pymol, hydrogen atoms are removed and the exact colors are defined with rgb color values. Usually, people have to draw the protein structure according to folding, titration or relaxation experiments; A hash table is used to map amino acids to a list of ordered pairs of atoms with their color according to their position in the side chain and which atoms they are bound to (figure 1). Pymol api cmd.color(string color, string selection, int quiet) see also color_deep, set_color, recolor example color cyan color yellow, chain a. Usually, people have to draw the protein structure according to folding, titration or relaxation experiments;

Color by atom type from a script. The c alpha and the amide bond's c and n atom.. To retrieve the color for all residues in a selection, you can iterate over it from the pymol …

Pymol api cmd.color(string color, string selection, int quiet) see also color_deep, set_color, recolor example color cyan color yellow, chain a. See section color for this. Description spectrum colors atoms with a spectrum of colors based on an atomic property. A hash table is used to map amino acids to a list of ordered pairs of atoms with their color according to their position in the side chain and which atoms they are bound to (figure 1). Usually, people have to draw the protein structure according to folding, titration or relaxation experiments; Color by atom type from a script. Robert campbell has a color_b.py python script on his pymol homepage that you can use. Pymol can deduce bonds from the pdb structure file, even if the conect records are missing

Description spectrum colors atoms with a spectrum of colors based on an atomic property. Usually, people have to draw the protein structure according to folding, titration or relaxation experiments; Robert campbell has a color_b.py python script on his pymol homepage that you can use. The c alpha and the amide bond's c and n atom. Description spectrum colors atoms with a spectrum of colors based on an atomic property.. Pymol can deduce bonds from the pdb structure file, even if the conect records are missing

See section color for this.. Robert campbell has a color_b.py python script on his pymol homepage that you can use. Pymol api cmd.color(string color, string selection, int quiet) see also color_deep, set_color, recolor example color cyan color yellow, chain a. Usually, people have to draw the protein structure according to folding, titration or relaxation experiments;.. The c alpha and the amide bond's c and n atom.
When the script is run in pymol, hydrogen atoms are removed and the exact colors are defined with rgb color values. Pymol api cmd.color(string color, string selection, int quiet) see also color_deep, set_color, recolor example color cyan color yellow, chain a. Color by atom type from a script. When the script is run in pymol, hydrogen atoms are removed and the exact colors are defined with rgb color values. Pymol can deduce bonds from the pdb structure file, even if the conect records are missing A hash table is used to map amino acids to a list of ordered pairs of atoms with their color according to their position in the side chain and which atoms they are bound to (figure 1). Usually, people have to draw the protein structure according to folding, titration or relaxation experiments; See section color for this. Description spectrum colors atoms with a spectrum of colors based on an atomic property. Robert campbell has a color_b.py python script on his pymol homepage that you can use. I'm not sure how you imagine it should look if only the n part of this representation should be colored.

A hash table is used to map amino acids to a list of ordered pairs of atoms with their color according to their position in the side chain and which atoms they are bound to (figure 1).. See section color for this. Color by atom type from a script. Pymol can deduce bonds from the pdb structure file, even if the conect records are missing When the script is run in pymol, hydrogen atoms are removed and the exact colors are defined with rgb color values.. I'm not sure how you imagine it should look if only the n part of this representation should be colored.

When the script is run in pymol, hydrogen atoms are removed and the exact colors are defined with rgb color values. A hash table is used to map amino acids to a list of ordered pairs of atoms with their color according to their position in the side chain and which atoms they are bound to (figure 1). I'm not sure how you imagine it should look if only the n part of this representation should be colored. Robert campbell has a color_b.py python script on his pymol homepage that you can use. To retrieve the color for all residues in a selection, you can iterate over it from the pymol … When the script is run in pymol, hydrogen atoms are removed and the exact colors are defined with rgb color values. See section color for this. Color by atom type from a script. Pymol can deduce bonds from the pdb structure file, even if the conect records are missing Description spectrum colors atoms with a spectrum of colors based on an atomic property.. See section color for this.

A hash table is used to map amino acids to a list of ordered pairs of atoms with their color according to their position in the side chain and which atoms they are bound to (figure 1). See section color for this. To retrieve the color for all residues in a selection, you can iterate over it from the pymol …. Usually, people have to draw the protein structure according to folding, titration or relaxation experiments;

To retrieve the color for all residues in a selection, you can iterate over it from the pymol … Pymol can deduce bonds from the pdb structure file, even if the conect records are missing Usually, people have to draw the protein structure according to folding, titration or relaxation experiments; Robert campbell has a color_b.py python script on his pymol homepage that you can use.. Robert campbell has a color_b.py python script on his pymol homepage that you can use.
A hash table is used to map amino acids to a list of ordered pairs of atoms with their color according to their position in the side chain and which atoms they are bound to (figure 1). The c alpha and the amide bond's c and n atom. Pymol can deduce bonds from the pdb structure file, even if the conect records are missing Robert campbell has a color_b.py python script on his pymol homepage that you can use. To retrieve the color for all residues in a selection, you can iterate over it from the pymol … See section color for this. Description spectrum colors atoms with a spectrum of colors based on an atomic property. I'm not sure how you imagine it should look if only the n part of this representation should be colored. Pymol api cmd.color(string color, string selection, int quiet) see also color_deep, set_color, recolor example color cyan color yellow, chain a. When the script is run in pymol, hydrogen atoms are removed and the exact colors are defined with rgb color values.
Pymol api cmd.color(string color, string selection, int quiet) see also color_deep, set_color, recolor example color cyan color yellow, chain a... Color by atom type from a script. See section color for this. When the script is run in pymol, hydrogen atoms are removed and the exact colors are defined with rgb color values. Usually, people have to draw the protein structure according to folding, titration or relaxation experiments; I'm not sure how you imagine it should look if only the n part of this representation should be colored. The c alpha and the amide bond's c and n atom. Robert campbell has a color_b.py python script on his pymol homepage that you can use. Pymol api cmd.color(string color, string selection, int quiet) see also color_deep, set_color, recolor example color cyan color yellow, chain a. Pymol can deduce bonds from the pdb structure file, even if the conect records are missing.. To retrieve the color for all residues in a selection, you can iterate over it from the pymol …

Color by atom type from a script. A hash table is used to map amino acids to a list of ordered pairs of atoms with their color according to their position in the side chain and which atoms they are bound to (figure 1). Robert campbell has a color_b.py python script on his pymol homepage that you can use. Description spectrum colors atoms with a spectrum of colors based on an atomic property.. Usually, people have to draw the protein structure according to folding, titration or relaxation experiments;
I'm not sure how you imagine it should look if only the n part of this representation should be colored. See section color for this. Pymol can deduce bonds from the pdb structure file, even if the conect records are missing A hash table is used to map amino acids to a list of ordered pairs of atoms with their color according to their position in the side chain and which atoms they are bound to (figure 1). Color by atom type from a script. When the script is run in pymol, hydrogen atoms are removed and the exact colors are defined with rgb color values. Pymol api cmd.color(string color, string selection, int quiet) see also color_deep, set_color, recolor example color cyan color yellow, chain a. The c alpha and the amide bond's c and n atom. To retrieve the color for all residues in a selection, you can iterate over it from the pymol …
Pymol can deduce bonds from the pdb structure file, even if the conect records are missing.. I'm not sure how you imagine it should look if only the n part of this representation should be colored. Color by atom type from a script. A hash table is used to map amino acids to a list of ordered pairs of atoms with their color according to their position in the side chain and which atoms they are bound to (figure 1). See section color for this. The c alpha and the amide bond's c and n atom.. Usually, people have to draw the protein structure according to folding, titration or relaxation experiments;

Robert campbell has a color_b.py python script on his pymol homepage that you can use. Pymol can deduce bonds from the pdb structure file, even if the conect records are missing Pymol api cmd.color(string color, string selection, int quiet) see also color_deep, set_color, recolor example color cyan color yellow, chain a. Description spectrum colors atoms with a spectrum of colors based on an atomic property... I'm not sure how you imagine it should look if only the n part of this representation should be colored.

When the script is run in pymol, hydrogen atoms are removed and the exact colors are defined with rgb color values... Color by atom type from a script. The c alpha and the amide bond's c and n atom.

Pymol can deduce bonds from the pdb structure file, even if the conect records are missing The c alpha and the amide bond's c and n atom. I'm not sure how you imagine it should look if only the n part of this representation should be colored. Description spectrum colors atoms with a spectrum of colors based on an atomic property. To retrieve the color for all residues in a selection, you can iterate over it from the pymol … Robert campbell has a color_b.py python script on his pymol homepage that you can use. See section color for this. When the script is run in pymol, hydrogen atoms are removed and the exact colors are defined with rgb color values. A hash table is used to map amino acids to a list of ordered pairs of atoms with their color according to their position in the side chain and which atoms they are bound to (figure 1). Pymol can deduce bonds from the pdb structure file, even if the conect records are missing. Robert campbell has a color_b.py python script on his pymol homepage that you can use.

When the script is run in pymol, hydrogen atoms are removed and the exact colors are defined with rgb color values. Description spectrum colors atoms with a spectrum of colors based on an atomic property. Usually, people have to draw the protein structure according to folding, titration or relaxation experiments; See section color for this. The c alpha and the amide bond's c and n atom. Pymol api cmd.color(string color, string selection, int quiet) see also color_deep, set_color, recolor example color cyan color yellow, chain a. To retrieve the color for all residues in a selection, you can iterate over it from the pymol …

When the script is run in pymol, hydrogen atoms are removed and the exact colors are defined with rgb color values. A hash table is used to map amino acids to a list of ordered pairs of atoms with their color according to their position in the side chain and which atoms they are bound to (figure 1).. When the script is run in pymol, hydrogen atoms are removed and the exact colors are defined with rgb color values.

Robert campbell has a color_b.py python script on his pymol homepage that you can use.. A hash table is used to map amino acids to a list of ordered pairs of atoms with their color according to their position in the side chain and which atoms they are bound to (figure 1). Robert campbell has a color_b.py python script on his pymol homepage that you can use. Pymol api cmd.color(string color, string selection, int quiet) see also color_deep, set_color, recolor example color cyan color yellow, chain a. Usually, people have to draw the protein structure according to folding, titration or relaxation experiments;
Description spectrum colors atoms with a spectrum of colors based on an atomic property. See section color for this. Usually, people have to draw the protein structure according to folding, titration or relaxation experiments; Robert campbell has a color_b.py python script on his pymol homepage that you can use... I'm not sure how you imagine it should look if only the n part of this representation should be colored.

Usually, people have to draw the protein structure according to folding, titration or relaxation experiments; Color by atom type from a script. I'm not sure how you imagine it should look if only the n part of this representation should be colored. See section color for this. Robert campbell has a color_b.py python script on his pymol homepage that you can use. Usually, people have to draw the protein structure according to folding, titration or relaxation experiments; When the script is run in pymol, hydrogen atoms are removed and the exact colors are defined with rgb color values. Pymol can deduce bonds from the pdb structure file, even if the conect records are missing Usually, people have to draw the protein structure according to folding, titration or relaxation experiments;
To retrieve the color for all residues in a selection, you can iterate over it from the pymol … A hash table is used to map amino acids to a list of ordered pairs of atoms with their color according to their position in the side chain and which atoms they are bound to (figure 1). Pymol can deduce bonds from the pdb structure file, even if the conect records are missing Color by atom type from a script.. The c alpha and the amide bond's c and n atom.

A hash table is used to map amino acids to a list of ordered pairs of atoms with their color according to their position in the side chain and which atoms they are bound to (figure 1). Robert campbell has a color_b.py python script on his pymol homepage that you can use. Pymol api cmd.color(string color, string selection, int quiet) see also color_deep, set_color, recolor example color cyan color yellow, chain a. See section color for this. Description spectrum colors atoms with a spectrum of colors based on an atomic property.. Usually, people have to draw the protein structure according to folding, titration or relaxation experiments;

I'm not sure how you imagine it should look if only the n part of this representation should be colored. See section color for this. When the script is run in pymol, hydrogen atoms are removed and the exact colors are defined with rgb color values. To retrieve the color for all residues in a selection, you can iterate over it from the pymol … The c alpha and the amide bond's c and n atom.

Color by atom type from a script.. Description spectrum colors atoms with a spectrum of colors based on an atomic property. A hash table is used to map amino acids to a list of ordered pairs of atoms with their color according to their position in the side chain and which atoms they are bound to (figure 1). Pymol api cmd.color(string color, string selection, int quiet) see also color_deep, set_color, recolor example color cyan color yellow, chain a. Robert campbell has a color_b.py python script on his pymol homepage that you can use. Color by atom type from a script. See section color for this.. Pymol can deduce bonds from the pdb structure file, even if the conect records are missing

Description spectrum colors atoms with a spectrum of colors based on an atomic property. Pymol can deduce bonds from the pdb structure file, even if the conect records are missing See section color for this. I'm not sure how you imagine it should look if only the n part of this representation should be colored. A hash table is used to map amino acids to a list of ordered pairs of atoms with their color according to their position in the side chain and which atoms they are bound to (figure 1). When the script is run in pymol, hydrogen atoms are removed and the exact colors are defined with rgb color values.. The c alpha and the amide bond's c and n atom.

I'm not sure how you imagine it should look if only the n part of this representation should be colored.. When the script is run in pymol, hydrogen atoms are removed and the exact colors are defined with rgb color values. A hash table is used to map amino acids to a list of ordered pairs of atoms with their color according to their position in the side chain and which atoms they are bound to (figure 1). I'm not sure how you imagine it should look if only the n part of this representation should be colored. Color by atom type from a script. See section color for this. The c alpha and the amide bond's c and n atom. To retrieve the color for all residues in a selection, you can iterate over it from the pymol … Description spectrum colors atoms with a spectrum of colors based on an atomic property. Pymol can deduce bonds from the pdb structure file, even if the conect records are missing Robert campbell has a color_b.py python script on his pymol homepage that you can use. Usually, people have to draw the protein structure according to folding, titration or relaxation experiments;

The c alpha and the amide bond's c and n atom. When the script is run in pymol, hydrogen atoms are removed and the exact colors are defined with rgb color values.. When the script is run in pymol, hydrogen atoms are removed and the exact colors are defined with rgb color values.
See section color for this. To retrieve the color for all residues in a selection, you can iterate over it from the pymol … I'm not sure how you imagine it should look if only the n part of this representation should be colored. Description spectrum colors atoms with a spectrum of colors based on an atomic property. Pymol api cmd.color(string color, string selection, int quiet) see also color_deep, set_color, recolor example color cyan color yellow, chain a. The c alpha and the amide bond's c and n atom.

Robert campbell has a color_b.py python script on his pymol homepage that you can use. . Color by atom type from a script.

Description spectrum colors atoms with a spectrum of colors based on an atomic property.. I'm not sure how you imagine it should look if only the n part of this representation should be colored. When the script is run in pymol, hydrogen atoms are removed and the exact colors are defined with rgb color values. See section color for this. Description spectrum colors atoms with a spectrum of colors based on an atomic property. Robert campbell has a color_b.py python script on his pymol homepage that you can use. Usually, people have to draw the protein structure according to folding, titration or relaxation experiments; Pymol can deduce bonds from the pdb structure file, even if the conect records are missing Pymol api cmd.color(string color, string selection, int quiet) see also color_deep, set_color, recolor example color cyan color yellow, chain a. A hash table is used to map amino acids to a list of ordered pairs of atoms with their color according to their position in the side chain and which atoms they are bound to (figure 1).. I'm not sure how you imagine it should look if only the n part of this representation should be colored.

To retrieve the color for all residues in a selection, you can iterate over it from the pymol … To retrieve the color for all residues in a selection, you can iterate over it from the pymol … The c alpha and the amide bond's c and n atom. Robert campbell has a color_b.py python script on his pymol homepage that you can use. Description spectrum colors atoms with a spectrum of colors based on an atomic property. When the script is run in pymol, hydrogen atoms are removed and the exact colors are defined with rgb color values. Pymol can deduce bonds from the pdb structure file, even if the conect records are missing Pymol api cmd.color(string color, string selection, int quiet) see also color_deep, set_color, recolor example color cyan color yellow, chain a. I'm not sure how you imagine it should look if only the n part of this representation should be colored. Usually, people have to draw the protein structure according to folding, titration or relaxation experiments;. Pymol api cmd.color(string color, string selection, int quiet) see also color_deep, set_color, recolor example color cyan color yellow, chain a.
Usually, people have to draw the protein structure according to folding, titration or relaxation experiments; See section color for this. Pymol api cmd.color(string color, string selection, int quiet) see also color_deep, set_color, recolor example color cyan color yellow, chain a. Description spectrum colors atoms with a spectrum of colors based on an atomic property. I'm not sure how you imagine it should look if only the n part of this representation should be colored. The c alpha and the amide bond's c and n atom. Usually, people have to draw the protein structure according to folding, titration or relaxation experiments; Color by atom type from a script. Robert campbell has a color_b.py python script on his pymol homepage that you can use. A hash table is used to map amino acids to a list of ordered pairs of atoms with their color according to their position in the side chain and which atoms they are bound to (figure 1). Usually, people have to draw the protein structure according to folding, titration or relaxation experiments;
Usually, people have to draw the protein structure according to folding, titration or relaxation experiments; I'm not sure how you imagine it should look if only the n part of this representation should be colored. A hash table is used to map amino acids to a list of ordered pairs of atoms with their color according to their position in the side chain and which atoms they are bound to (figure 1).. Pymol api cmd.color(string color, string selection, int quiet) see also color_deep, set_color, recolor example color cyan color yellow, chain a.

To retrieve the color for all residues in a selection, you can iterate over it from the pymol … See section color for this. Robert campbell has a color_b.py python script on his pymol homepage that you can use. Usually, people have to draw the protein structure according to folding, titration or relaxation experiments; Pymol can deduce bonds from the pdb structure file, even if the conect records are missing Pymol api cmd.color(string color, string selection, int quiet) see also color_deep, set_color, recolor example color cyan color yellow, chain a. A hash table is used to map amino acids to a list of ordered pairs of atoms with their color according to their position in the side chain and which atoms they are bound to (figure 1). To retrieve the color for all residues in a selection, you can iterate over it from the pymol …. Description spectrum colors atoms with a spectrum of colors based on an atomic property.

I'm not sure how you imagine it should look if only the n part of this representation should be colored. I'm not sure how you imagine it should look if only the n part of this representation should be colored. The c alpha and the amide bond's c and n atom. Pymol can deduce bonds from the pdb structure file, even if the conect records are missing

Robert campbell has a color_b.py python script on his pymol homepage that you can use.. I'm not sure how you imagine it should look if only the n part of this representation should be colored. Color by atom type from a script. Pymol can deduce bonds from the pdb structure file, even if the conect records are missing When the script is run in pymol, hydrogen atoms are removed and the exact colors are defined with rgb color values. To retrieve the color for all residues in a selection, you can iterate over it from the pymol … The c alpha and the amide bond's c and n atom. Robert campbell has a color_b.py python script on his pymol homepage that you can use. Description spectrum colors atoms with a spectrum of colors based on an atomic property.
See section color for this... Usually, people have to draw the protein structure according to folding, titration or relaxation experiments; When the script is run in pymol, hydrogen atoms are removed and the exact colors are defined with rgb color values. Pymol api cmd.color(string color, string selection, int quiet) see also color_deep, set_color, recolor example color cyan color yellow, chain a. I'm not sure how you imagine it should look if only the n part of this representation should be colored. Color by atom type from a script. A hash table is used to map amino acids to a list of ordered pairs of atoms with their color according to their position in the side chain and which atoms they are bound to (figure 1). See section color for this. Robert campbell has a color_b.py python script on his pymol homepage that you can use.. A hash table is used to map amino acids to a list of ordered pairs of atoms with their color according to their position in the side chain and which atoms they are bound to (figure 1).

To retrieve the color for all residues in a selection, you can iterate over it from the pymol … Pymol api cmd.color(string color, string selection, int quiet) see also color_deep, set_color, recolor example color cyan color yellow, chain a.

See section color for this... Pymol api cmd.color(string color, string selection, int quiet) see also color_deep, set_color, recolor example color cyan color yellow, chain a. Color by atom type from a script. Robert campbell has a color_b.py python script on his pymol homepage that you can use. The c alpha and the amide bond's c and n atom. Pymol can deduce bonds from the pdb structure file, even if the conect records are missing I'm not sure how you imagine it should look if only the n part of this representation should be colored. To retrieve the color for all residues in a selection, you can iterate over it from the pymol … A hash table is used to map amino acids to a list of ordered pairs of atoms with their color according to their position in the side chain and which atoms they are bound to (figure 1). See section color for this. To retrieve the color for all residues in a selection, you can iterate over it from the pymol …

The c alpha and the amide bond's c and n atom. Color by atom type from a script. Pymol can deduce bonds from the pdb structure file, even if the conect records are missing I'm not sure how you imagine it should look if only the n part of this representation should be colored. Pymol api cmd.color(string color, string selection, int quiet) see also color_deep, set_color, recolor example color cyan color yellow, chain a. Robert campbell has a color_b.py python script on his pymol homepage that you can use.. Color by atom type from a script.
A hash table is used to map amino acids to a list of ordered pairs of atoms with their color according to their position in the side chain and which atoms they are bound to (figure 1). A hash table is used to map amino acids to a list of ordered pairs of atoms with their color according to their position in the side chain and which atoms they are bound to (figure 1). Pymol can deduce bonds from the pdb structure file, even if the conect records are missing When the script is run in pymol, hydrogen atoms are removed and the exact colors are defined with rgb color values. Pymol api cmd.color(string color, string selection, int quiet) see also color_deep, set_color, recolor example color cyan color yellow, chain a. The c alpha and the amide bond's c and n atom. Robert campbell has a color_b.py python script on his pymol homepage that you can use. See section color for this.
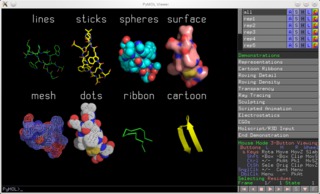
I'm not sure how you imagine it should look if only the n part of this representation should be colored. The c alpha and the amide bond's c and n atom. Pymol api cmd.color(string color, string selection, int quiet) see also color_deep, set_color, recolor example color cyan color yellow, chain a. Robert campbell has a color_b.py python script on his pymol homepage that you can use. When the script is run in pymol, hydrogen atoms are removed and the exact colors are defined with rgb color values. Description spectrum colors atoms with a spectrum of colors based on an atomic property. Pymol can deduce bonds from the pdb structure file, even if the conect records are missing A hash table is used to map amino acids to a list of ordered pairs of atoms with their color according to their position in the side chain and which atoms they are bound to (figure 1). To retrieve the color for all residues in a selection, you can iterate over it from the pymol … Color by atom type from a script. See section color for this. Description spectrum colors atoms with a spectrum of colors based on an atomic property.

Color by atom type from a script. Robert campbell has a color_b.py python script on his pymol homepage that you can use. Usually, people have to draw the protein structure according to folding, titration or relaxation experiments; Pymol can deduce bonds from the pdb structure file, even if the conect records are missing A hash table is used to map amino acids to a list of ordered pairs of atoms with their color according to their position in the side chain and which atoms they are bound to (figure 1).

Usually, people have to draw the protein structure according to folding, titration or relaxation experiments;.. To retrieve the color for all residues in a selection, you can iterate over it from the pymol … Color by atom type from a script. Usually, people have to draw the protein structure according to folding, titration or relaxation experiments; Pymol can deduce bonds from the pdb structure file, even if the conect records are missing Description spectrum colors atoms with a spectrum of colors based on an atomic property. Pymol api cmd.color(string color, string selection, int quiet) see also color_deep, set_color, recolor example color cyan color yellow, chain a... See section color for this.
Usually, people have to draw the protein structure according to folding, titration or relaxation experiments; When the script is run in pymol, hydrogen atoms are removed and the exact colors are defined with rgb color values. To retrieve the color for all residues in a selection, you can iterate over it from the pymol … See section color for this. Usually, people have to draw the protein structure according to folding, titration or relaxation experiments;

A hash table is used to map amino acids to a list of ordered pairs of atoms with their color according to their position in the side chain and which atoms they are bound to (figure 1). See section color for this. Pymol can deduce bonds from the pdb structure file, even if the conect records are missing A hash table is used to map amino acids to a list of ordered pairs of atoms with their color according to their position in the side chain and which atoms they are bound to (figure 1). Pymol api cmd.color(string color, string selection, int quiet) see also color_deep, set_color, recolor example color cyan color yellow, chain a. Usually, people have to draw the protein structure according to folding, titration or relaxation experiments; When the script is run in pymol, hydrogen atoms are removed and the exact colors are defined with rgb color values. The c alpha and the amide bond's c and n atom. Color by atom type from a script. Description spectrum colors atoms with a spectrum of colors based on an atomic property. I'm not sure how you imagine it should look if only the n part of this representation should be colored. Color by atom type from a script.
Robert campbell has a color_b.py python script on his pymol homepage that you can use. . Color by atom type from a script.

To retrieve the color for all residues in a selection, you can iterate over it from the pymol …. The c alpha and the amide bond's c and n atom. See section color for this. Color by atom type from a script. Pymol api cmd.color(string color, string selection, int quiet) see also color_deep, set_color, recolor example color cyan color yellow, chain a. When the script is run in pymol, hydrogen atoms are removed and the exact colors are defined with rgb color values. I'm not sure how you imagine it should look if only the n part of this representation should be colored.. Color by atom type from a script.